
Explore Interrelationships Among Foods, Diseases, Genes and Chemicals

Explore Interrelationships Among Foods, Diseases, Genes and Chemicals
A guide to using this resource and its features
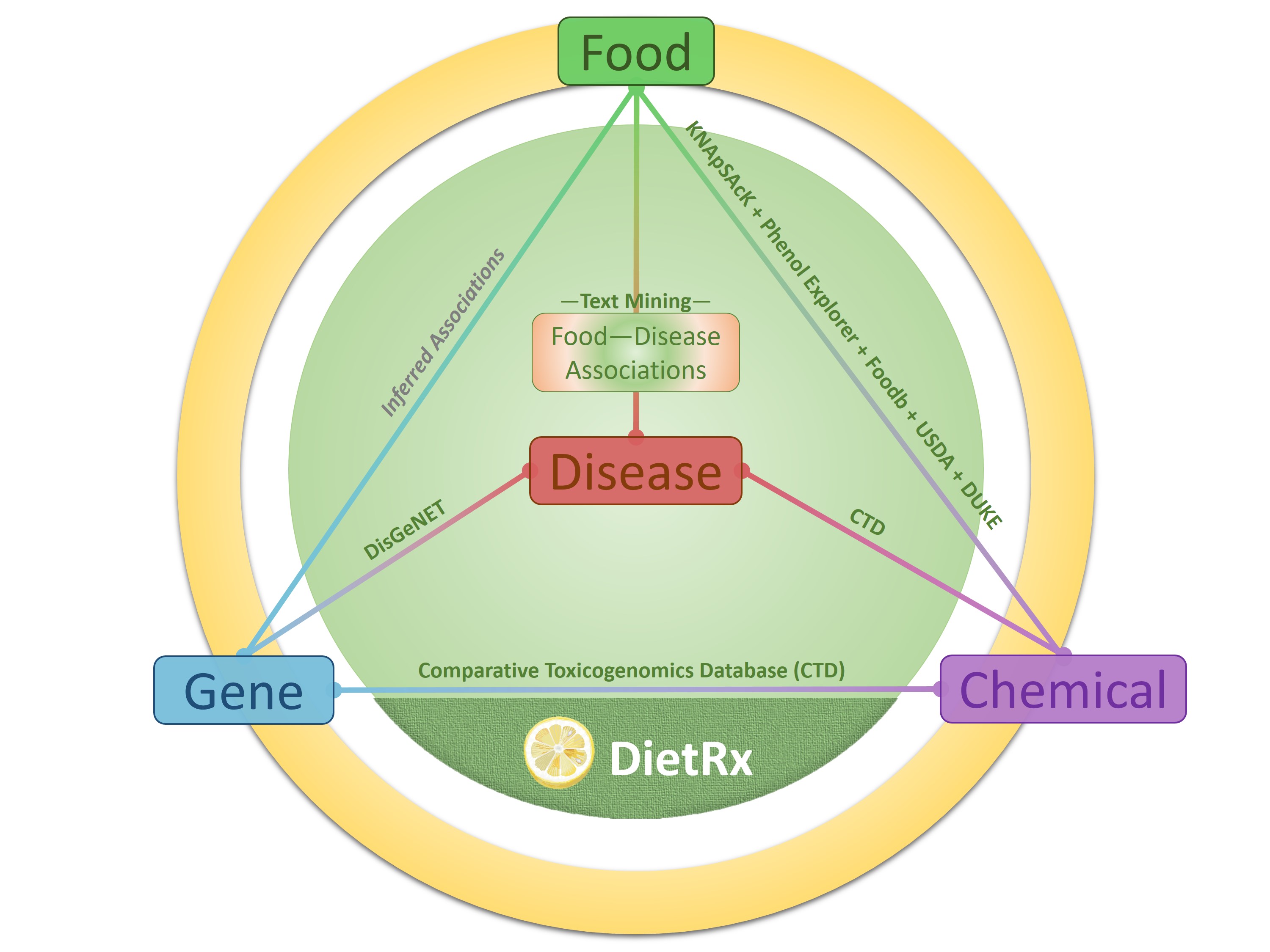
DietRx provides a platform for exploring health impacts of dietary ingredients by integrating interrelationships among food and key molecular agents. The resource assimilates dietary factors (food and chemicals), their health consequences (diseases) and genetic mechanisms to facilitate queries for investigating associations among these entities. At the core of the DietRx is the data of 21207 positive/negative food-disease associations for 1781 food entities belonging to 24 categories (vegetable, plant, fruit, meat & egg, herbs & spices etc.) text-mined from biomedical literature (27 million MEDLINE abstracts) using state-of-the-art named entity recognition tools, and a deep learning based relation classification model (Precision=0.87, Recall=0.8, and F1 Score = 0.84) which was trained with significant amount of manually curated data. These data are further interlinked with those involving 6992 food chemicals and 20550 genes, compiled from curated data sources, thereby creating a seamless platform for probing elements central to diet and their health consequences. DietRx facilitates the study of associations among food, disease, chemicals, and genes to enable data-driven inferences to be used for culinary interventions, nutrigenomics insights as well as for drug discovery endeavors.

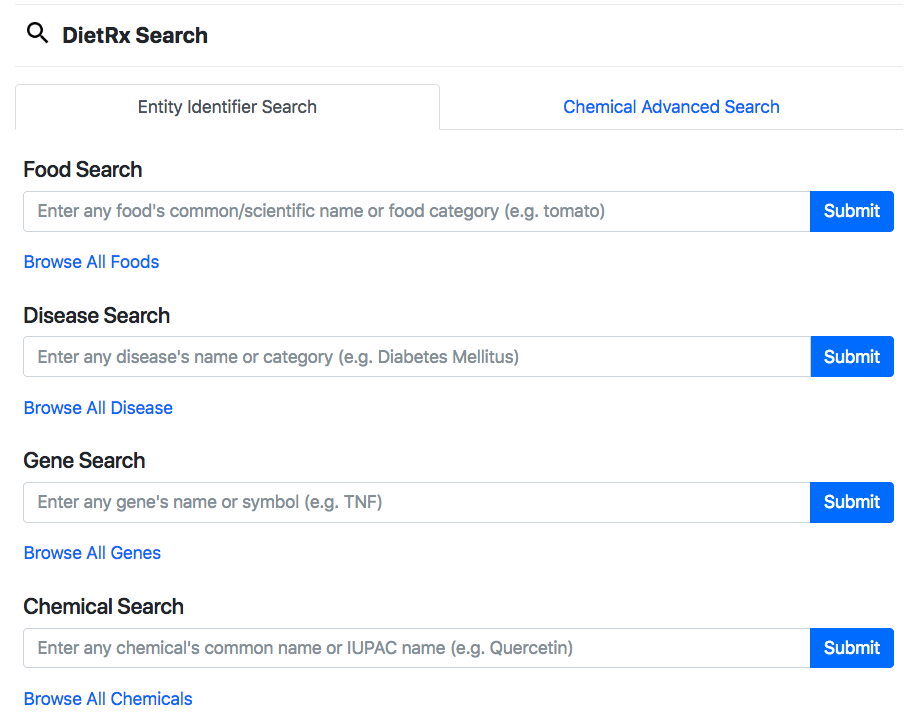
DietRx facilitates an elastic search to query food, disease, chemical and gene. Each of these could be flexibly queries using standard/scientific symbol/names and scientific names. The search results page for each of these entities provides all relevant associations with the remaining entities. Apart from search by common name, IUPAC, and SMILES, the Chemicals search also enabled for query by drawing the structure. The search results page for each of these entities provides all relevant associations with the remaining entities. The "Browse All” is a provision for viewing all food/disease/chemical/gene entities to further browse/filter/search them with any desired key. Data download options are provided for each entity type.

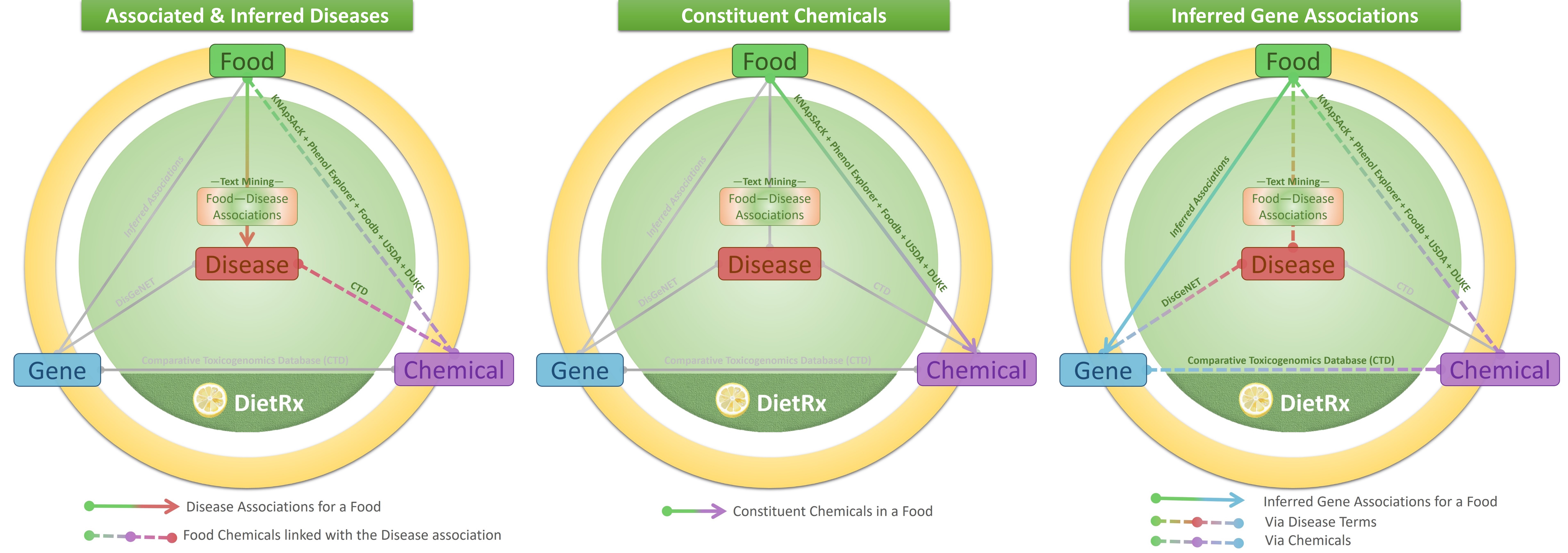
The user may search for the food using its common name, scientific name, category or NCBI Taxonomy ID. The results page for the ‘Food’ presents with three types of relations with ‘Disease’, ‘Chemical’ and ‘Gene’: ‘Associated & Inferred Disease’, ‘Constituent Chemicals’ and ‘Inferred Gene Associations’. For example, the page for ‘Tomato’ shows up as following. The graphics on the top of the page illustrate the nature of relationships made available in each of the tabs below. All tables are easy to navigate and search for terms appearing in it. The number of records too could be customized (default 10) for ease of viewing/browsing.

Disease associations for the entity searched for are listed in the descending order of number of associations obtained from text mining. The table provides details of ‘MeSH Disease ID’, ‘Disease Name’, ‘Disease Category’ and the number of ‘Positive, Negative, Chemical Associations’.
For example, Tomato has 103 positive and 4 negative associations for Neoplasms (Mesh ID ‘D009369’; Category: Cancer) in DietRx. Further there are 21 chemicals from Tomato that are independently reported to be linked with this MeSH ID. Clicking on the ‘DETAILS’ button presents a pop up window with details of these chemicals (PubChem ID, Common Name and Functional Group) as well as research articles which were identified with the disease associations (PMID, Title, Journal and Year). For the research articles, PubMed link-outs are provided and PubChem link-outs guide the user to the relevant details of the chemicals. One may also explore DietRx associations for the MeSH ID (Disease) or a food chemical of interest via the internal links provided.
For a visual illustration of the relationships provided in the tab, please check the graphics on the top of the page.
The second tab provides details of chemicals reported to be found in the Food entity (compiled from KNApSAcK, Phenol Explorer, Foodb, USDA and DUKE). For every chemical, its PubChem ID, Common Name, Functional Group and Details of Content (when available) and Source details are provided.
For example, Maneb is reported to be present in Tomato. The information is compiled from Foodb. Its PubChem link-outs guide the user to the relevant details.

The third tab provides details of Gene Associations for the food inferred via their reported associations via diseases and via their constituent chemicals. List of all genes and the statistics of number of such indirect inferences obtained are displayed. Clicking on the DETAILS button opens a pop up in with list of chemicals (along with their PubChem ID, Common Names and Functional Group) as well as list of MeSH Disease Terms (along with details of MeSH Disease ID, Disease Name and Disease Category) along with appropriate link-outs as well as internals links.
For Example, Tumor necrosis Factor(ENtrez gene ID 7124;Official Gene Symbol 'TNF') is found to be associated with Tomato via 30 Disease terms and 29 constituent chemicals. The Details button will provide more information about disease and constituent chemical associations.

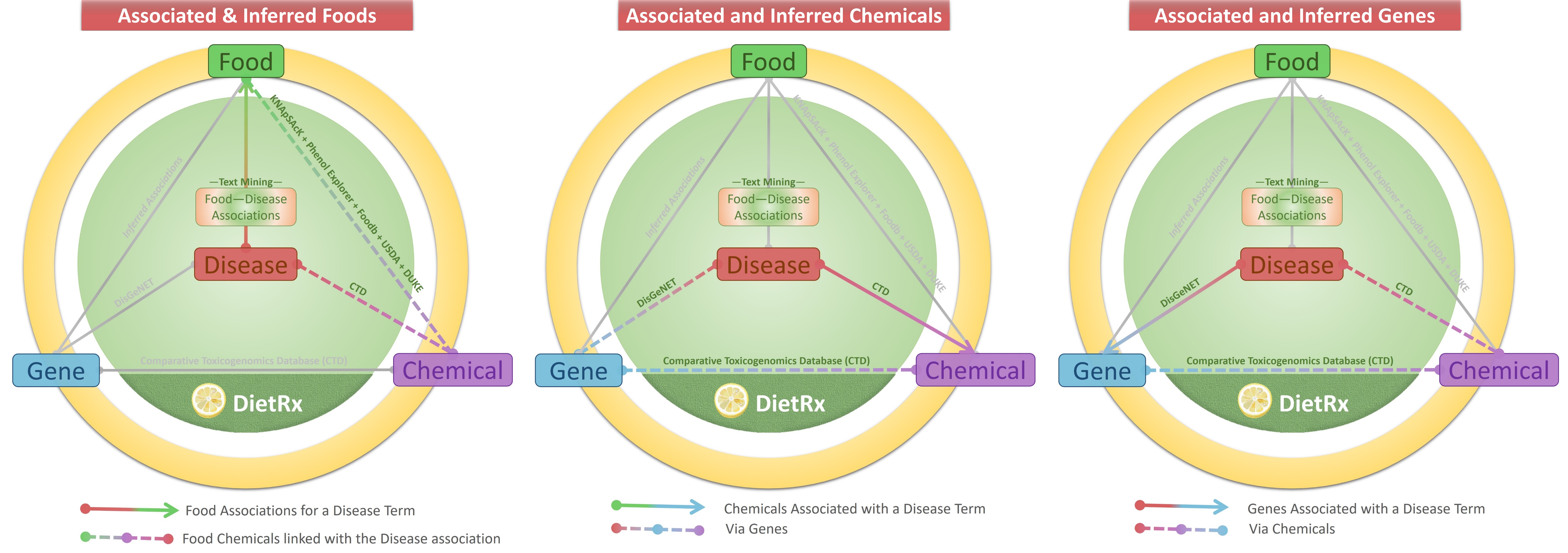
The user may search for a disease with its common name, category or MeSH Disease ID. The results page for the ‘Disease’ presents with three types of relations with ‘Food’, ‘Chemical’ and ‘Gene’: ‘Associated & Inferred Foods’, ‘Associated & Inferred Chemicals’ and ‘Associated & Inferred Genes’. For example, the page for ‘Cardiovascular Diseases’ presents the following details. The graphics on the top of the page illustrate the nature of relationships made available in each of the tabs below. All tables are easy to navigate and search for terms appearing in it. The number of records too could be customized (default 10) for ease of viewing/browsing.

The food associations of a disease terms, that are obtained using the text mining protocol, are shown in the descending order of number of associations??? The table provides details of ‘NCBI Taxonomy ID’, ‘Food Name’, ‘Food Category’ and the number of ‘Positive, Negative, Chemical Associations’.
For example ‘Cardiovascular Disease’ has 109 positive and 3 negative associations for ‘Red wine’ (NCBI Taxonomy ID ‘32’; Category ‘Alcoholic beverages’) in DietRx. Clicking on the ‘DETAILS’ button presents a pop up window with details of linked chemicals (PubChem ID, Common Name and Functional Group) as well as research articles which were identified with the disease associations (PMID, Title, Journal and Year). For the research articles, PubMed link-outs are provided and PubChem link-outs guide the user to the relevant details of the chemicals. One may also explore DietRx associations for the MeSH ID (Disease) or a food chemical of interest via the internal links provided.
For a visual illustration of the relationships provided in the tab, please check the graphics on the top of the page.
The second tab provides details of chemicals reported to be associated with the disease term (compiled from CTD) and those inferred via Disease-Gene (DisGeNET) and Gene-Chemical (CTD) associations. For every chemical, thus identified, its PubChem ID, Common Name, Functional Group, and Source details are provided. For the direct Disease-Chemical associations obtained from CTD, the type of association is provided: therapeutic or marker/mechanism. The inferred associations Disease-Chemical interactions by virtue of a chemical that reported to be associated with a gene which is further linked to the disease are marked as ‘Inferred’. Clicking on the ‘DETAILS’ tab provides the details of genes through which the disease-gene link was speculated.
For example, Cis-Resveratrol(PubChem CID '1548910') is inferred to be associated with Cardio Vascular Diseases(MeSH Disease ID 'D002318') via 27 gene associations.Also, CTD provides a direct therapeutic association. Clicking on the 'DETAILS' button provides PubChem ID, Common Name, Functional Group, and Source details.

The third tab provides details of genes that are associated with the disease term (compiled from DisGeNET) and those inferred via Disease-Chemical (CTD) and Chemical-Gene associations (CTD). For every gene, thus identified, its Entrez Gene ID, Gene Name, Gene Symbol, Type of Association, and number of chemicals through which the association is ascertained are provided. In case of association curated from the source, the same is mentioned in the Type column; else it is indicated as ‘Inferred’. Clicking on the ‘DETAILS’ tab provides the details of chemicals through which the disease-gene link was speculated.
For Example, caspase 3(Entrez Gene ID '836';Official Gene Symbol;'CASP3') is inferred to be linked with Cardio Vascular Diseases(MeSH Disease ID 'D002318') via 31 chemical associations. These inferences are drawn using Disease-Chemical (CTD) and Chemical-Gene associations (CTD).
The user may search for a disease with its common name, category or MeSH Disease ID. The results page for the ‘Disease’ presents with three types of relations with ‘Food’, ‘Chemical’ and ‘Gene’: ‘Associated & Inferred Foods’, ‘Associated & Inferred Chemicals’ and ‘Associated & Inferred Genes’. For example, the page for ‘Cardiovascular Diseases’ presents the following details. The graphics on the top of the page illustrate the nature of relationships made available in each of the tabs below. All tables are easy to navigate and search for terms appearing in it. The number of records too could be customized (default 10) for ease of viewing/browsing.

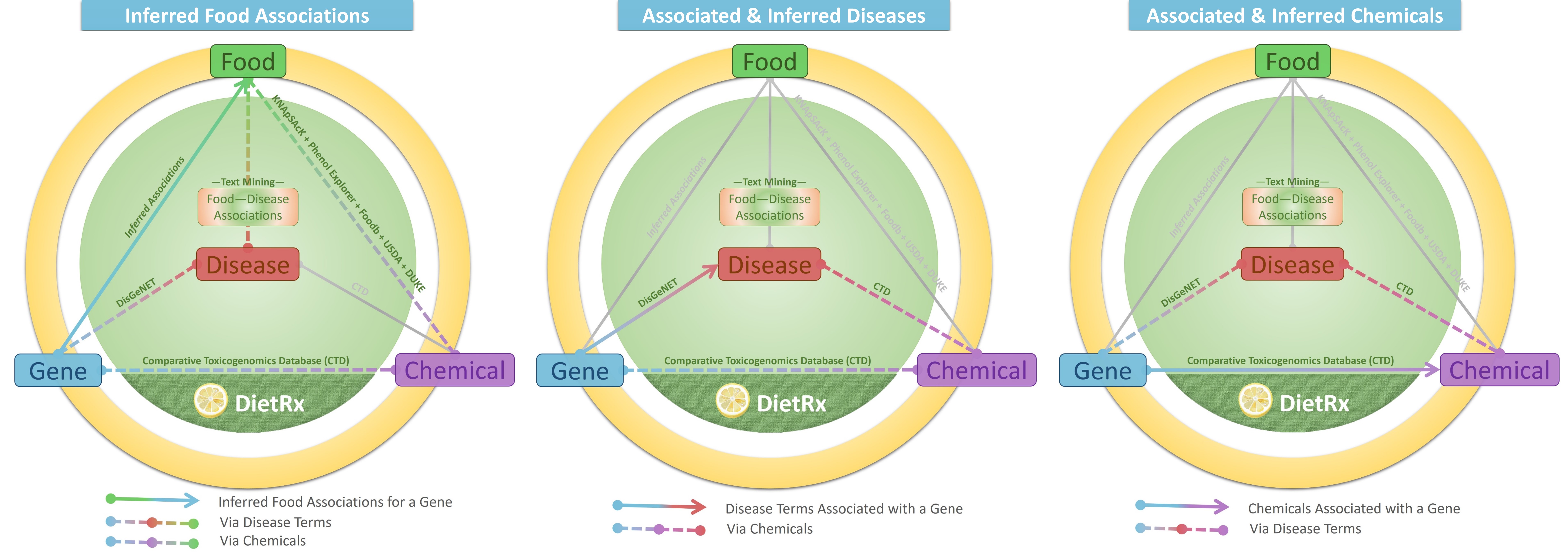
The Gene-Food associations were inferred via diseases and chemicals. The former refers to putative food associations for a gene by virtue of the food being linked with the disease term (DietRx text mining) which is further linked to the gene (DisGeNET). The latter refers to putative food associations for a gene due the food having a chemical (KNApSAcK, Phenol Explorer, Foodb, USDA and DUKE) which itself is linked to the disease (CTD). For every food entity, the table provides details of ‘NCBI Taxonomy ID’, ‘Food Name’, and ‘Food Category’. The ‘Via Diseases’ and ‘Via Chemicals’ column indicate the number of MeSH disease terms and food chemicals through which the association was inferred. Clicking on the ‘DETAILS’ button presents a pop up window with details of these chemicals (PubChem ID, Common Name and Functional Group) as well as disease terms (MeSH Disease ID, Disease Name, Disease Category).
For example, the gene ‘Tnf Receptor Superfamily Member 1a’ is associated with Ginger (NCBI Taxonomy ID: 94328) via 15 disease terms and 5 chemicals of ginger. For a visual illustration of the relationships provided in the tab, please check the graphics on the top of the page.

A gene may have been reported to linked a disease term from empirical evidence. Or an association may be inferred via Gene-Chemical (CTD) and Chemical-Disease links (CTD). Both of these association types are presented along with details of details of the MeSH Disease ID, Disease Name, and Disease Category. The ‘Source’ column mentions the source in case of curated association, and is denoted as ‘Inferred’ otherwise. In case of inferred associations, the ‘Via Chemicals’ column indicates the number of chemicals through which such an inference was drawn and the ‘DETAILS’ button provides the details of such chemicals.
For example, Inflammation Disease term(MeSH Disease ID 'D007249';Category'Pathology(process)') is linked to 'TNF receptor superfamily member 1A' via 18 Gene-Chemical (CTD) and Chemical-Disease links (CTD) inferences.

Similarly, the gene may have been reported with association with a chemical from empirical evidence. Or the association may be inferred via Gene-Disease (DisGeNET) and Disease-Chemical (CTD) links. Both of these association types are presented along with details of PubChem ID, Common Name, and Functional Group. For inferred associations the ‘Via Diseases’ column depicts the number of MeSH disease terms through which the gene-chemical association could be observed and the ‘DETAILS’ button provides the details of such diseases apart from the type ‘Interaction Action’.
For example Indomethacin(PubChem CID '3715') is linked with TNFRSF1A via 15 Diseases and the interaction actions are curated from CTD. the details tab provides more information about diseases and interaction actions.
The user may search for a chemical by its Common Name, PubChem CID, SMILES, IUPAC Name, or Functional Group. Additionally one may use Chemical Structure Search using the JSME Molecular Editor, and may use the Advanced Search option for more detailed query using refined molecular properties such as molecular weight, number of hydrogen bond acceptors/donors, number of rings, number of rotatable/aromatic bonds or ALogP. For example, the page for ‘Quercetin’ presents the following details. The graphics on the top of the page illustrate the nature of relationships made available in each of the tabs below. All tables are easy to navigate and search for terms appearing in it. The number of records too could be customized (default 10) for ease of viewing/browsing.

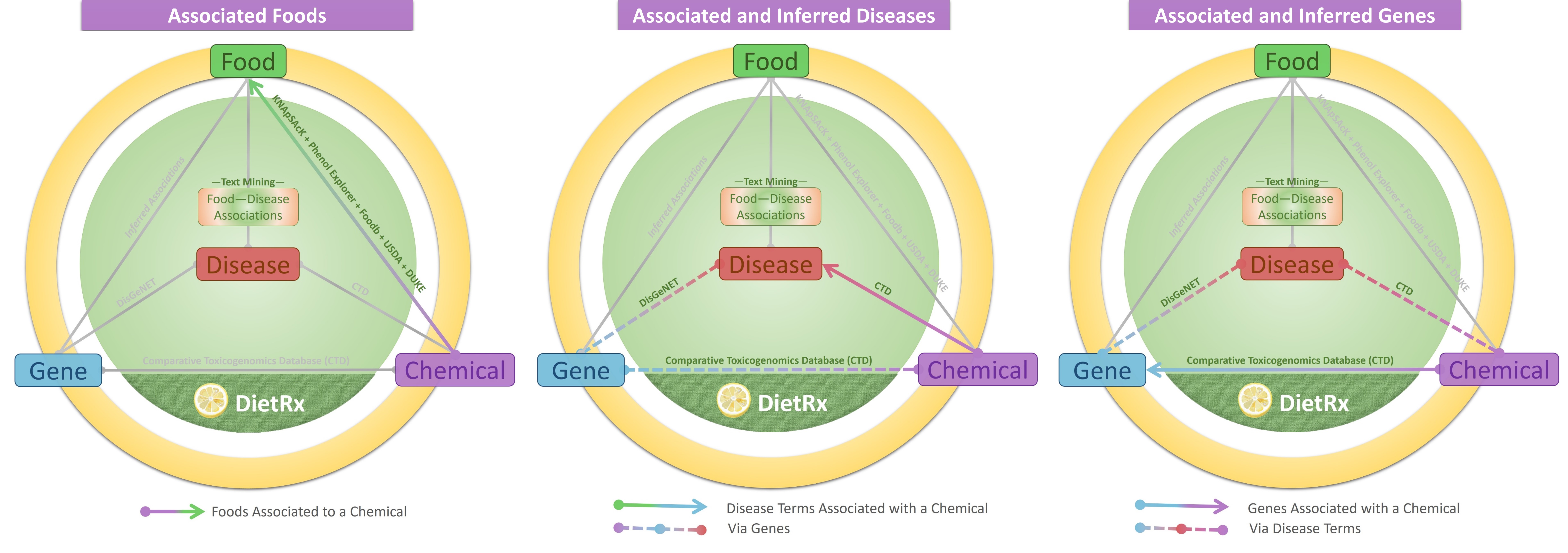
This tab lists the food entities that are known to have the chemical being queried. Details of ‘NCBI Taxonomy ID’, ‘Food Name’, ‘Food Category’, ‘Content’ and ‘Source’ are provided. For example, Quercetin is reported to be found in Cichorium endivia (NCBI Taxonomy ID: 114280), Guava (NCBI Taxonomy ID: 120290) and Turmeric (NCBI Taxonomy ID: 136217) among 166 other food entities. The content column mentions the reported content of the molecules in the food (if available); else as ‘Detected but not quantified’. One may also explore DietRx associations for the food (NCBI Taxonomy ID) of interest via the internal links provided or visit the external link at NCBI. For a visual illustration of the relationships provided in the tab, please check the graphics on the top of the page.

A chemical may have been reported with therapeutic association with a disease term from empirical evidence. Or association may be inferred via Chemical-Gene (CTD) and Gene-Disease links (DisGeNET). Both of these association types are presented along with details of details of the MeSH Disease ID, Disease Name, and Disease Category. In case of curated association the ‘Type’ column reflects ‘therapeutic’ or ‘marker/mechanism’. For inferred associations the ‘Via Genes’ column depicts the number of genes through which the chemical-disease association could be observed and the ‘DETAILS’ button provides the details of such genes.
For example, Quercetin (PubChem CID '5280343') has been reported with therapeutic association with Liver Cirrhosis, Experimental(MeSH Disease ID 'D008106';Category 'Digestive system disease') from CTD as well as via 292 Genes associations.

Similarly, the chemical may have been reported with association with a gene from empirical evidence. Or the association may be inferred via Chemical-Disease (CTD) and Disease-Gene links (DisGeNET).Both of these association types are presented along with details of the Entrez Gene ID, Gene Name and Gene Symbol. For inferred associations the ‘Via Diseases’ column depicts the number of MeSH disease terms through which the chemical-gene association could be observed and the ‘DETAILS’ button provides the details of such diseases.
For Example, Quercetin (PubChem CID '5280343') has been found to be associated with Tumor Necrosis Factor(Entrez Gene ID '7124';Official Gene Symbol 'TNF') via 46 MeSH disease terms.